Infrastructures in Fink
infrastructure
Fink infrastructure
Let’s take the opportunity of the bad turn of recent events to talk and learn a bit more on the infrastructure in place for Fink.
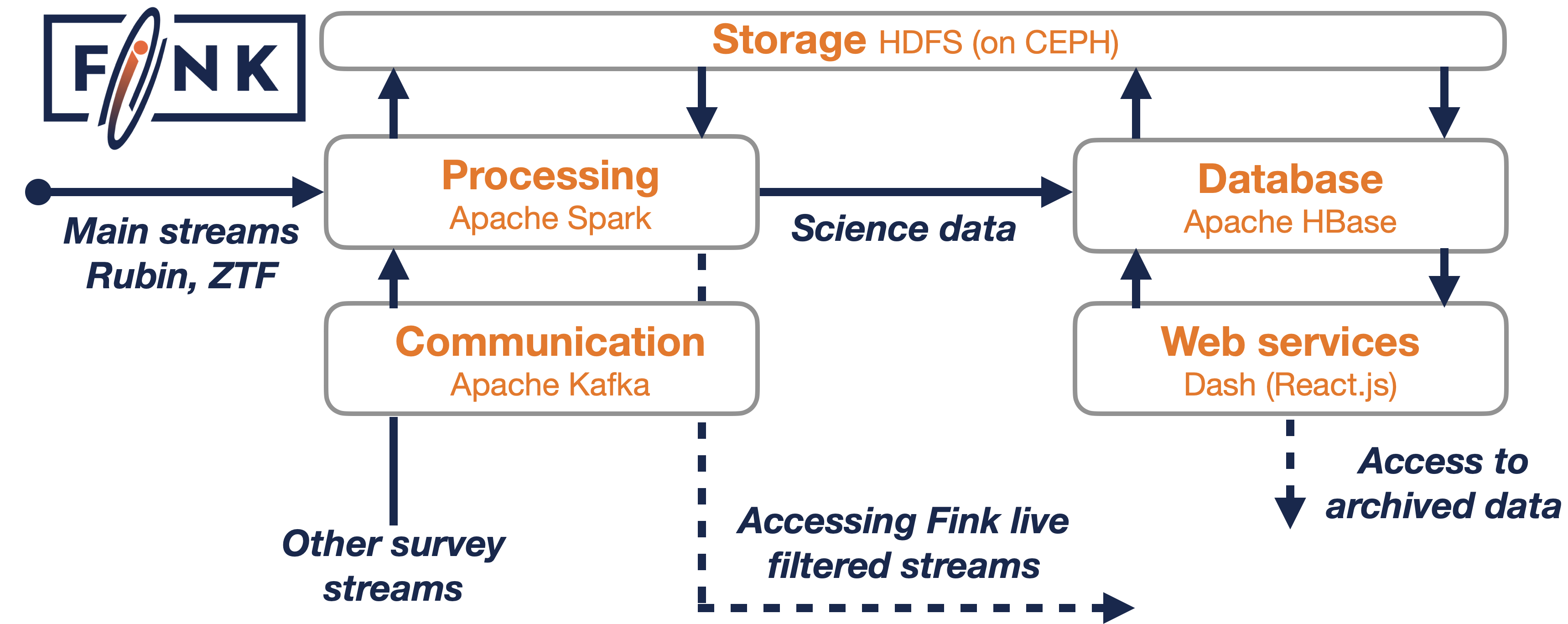 Fink is deployed on the cloud, that is each box in the Figure is an independent set of machines where specific software are running and executing tasks.
Fink is deployed on the cloud, that is each box in the Figure is an independent set of machines where specific software are running and executing tasks.
Nightly alert processing
The central part of the processing in Fink is the Apache Spark cluster (see fink-broker). It is currently a set of 12 machines (1 driver node, and 11 worker nodes). Each machine has 16 cores with 2 GB RAM per core. All machines use CentOs7 and we are currently running Apache Spark 2.4.7. Jobs are managed by Mesos, but we can also use Kubernetes since November 2021 (see fink-k8s).
ZTF, and Rubin in the future, is using Apache Kafka to produce a stream of alerts. The data is serialised using the Avro format. In Fink, we use the Spark-Kafka connector to directly poll alerts from the alert system. By using Apache Spark, we guarantee a very high throughput (tested nearly up to 10 times LSST rates, that is 300 MB/s over a long period) with a minimum of effort. Each night, we have a process waiting for the first alert of the night to come, and start Fink operations (otherwise we keep sleeping).
Alert data is polled and decoded on-the-fly upon receival. We make a copy of it on the Hadoop Distributed File System (HDFS), for reprocessing purposes. This stored data is not critical as it can be recomputed on-demand using the alert archive from the input telescope (we keep it for convenience for the moment).
Once alerts enter the Fink pipeline, boguses are rejected, and valid alerts are processed by the Fink science modules. Science modules are python module to extract additional information from the alerts (see fink-science). The processing is parallelised by Apache Spark to increase the throughput. The output of the processing is the incoming stream enriched with additional information, and it is stored on HDFS (temporary data, for later inspection purposes).
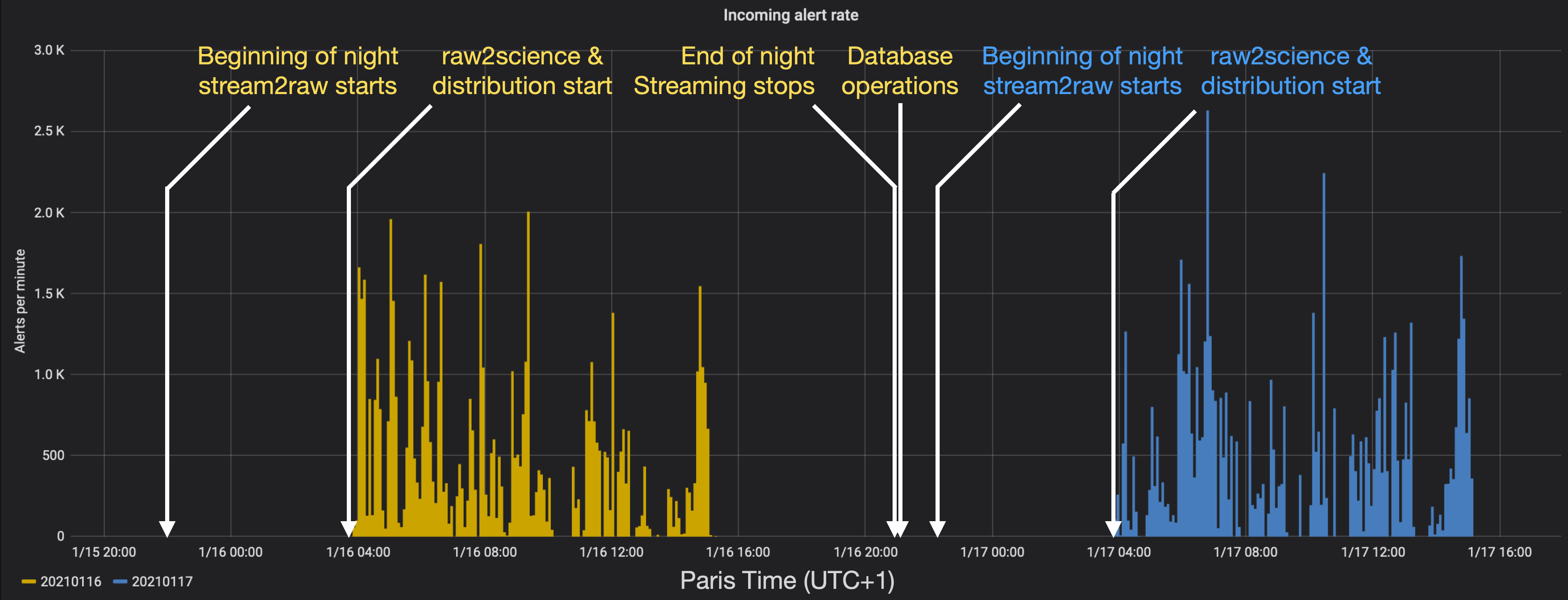 Fink operations as seen from the Grafana dashboard for the nights 20210116 and 20210117.
Fink operations as seen from the Grafana dashboard for the nights 20210116 and 20210117.
Storage system
Our main storage system is the Hadoop Distributed File System (HDFS). HDFS is a distributed file-system, providing very high aggregate bandwidth across the cluster. We currently have a cluster of 12 machines (1 namenode, and 11 datanodes), for a total of 45TB storage (with replication 3, that is 135TB of storage).
The good functioning of HDFS, as any distributed system, can be quite challenging for an intrinsically serial human brain. The error we are facing after the last power outage concerns HDFS, and the way it handles its metadata.
Real-time redirection
Right after the processing, we filter alerts based on their properties, and redirect alerts of interest to the community. Alerts are first redirected to our Apache Kafka cluster. This cluster is made of 5 machines, with 4 cores each, and 2 GB RAM for each core.
Each type of filter gives rise to a topic in Kafka. The filters are user-based, and can use anything inside alerts (see fink-filter). Users can then get the alerts in real-time using the Fink Kafka client (see fink-client).
In Kafka, streams are produced (and not sent), that is they are made available to the consumers that need to poll them to get the data (we do not send data automatically on the network). Alerts are available 4 days after their creation, so that users can get them on a later time if need be (or poll them again).
This cluster also handles alert simulations, by replaying ZTF streams (or any streams) at a tunable rate.
Science Portal & Database
We use Apache HBase to store our aggregated alert data (key-value storing). Apache HBase is a non-relational distributed database that allows to easily keep track of changes, to perform fast queries, and it allows flexibility in the data model schema. An HBase cluster has been deployed in the VirtualData cloud, and data is stored on HDFS (hence its unavailability when HDFS is dead). Our HBase instance is a single machine with 6 cores. The system currently uses an HBase-Spark connector to ingest and update data at the end of the night.
At the end of the observing night, we send all the processed alert data to our databases. There are many database operations:
- consolidate nightly temportary data on HDFS (optimising storage space, as alerts are tiny files stored on-the-fly on disks!)
- update the main HBase table with nightly data
- update the HBase index tables with nightly data
- send new early SN Ia candidates to the Transient Name server for follow-up
- perform aggregation operations, and update statistics
For ZTF, the whole processing takes about 15 minutes, on a dozen of cores.
We are managing different HBase tables serving different purposes: web application, light-curve post-processing and graph processing. I recently made a review talk on the use of HBase for Fink. Today, the main usage is for the Science Portal. Everytime you are searching something in the portal, or using directly the REST API, you are making queries inside HBase tables! They currently contain the data of about 40 million alerts (4TB).
The Science Portal is a web application deployed on a machine in the cloud with 8 cores. The application has been written with Dash (Python), a framework highly recommended! Internally, it uses the REST API to commmunicate with the database, and we also developped a custom HBase client in Java to efficiently make queries.